Nature Medicine - AI Section⭐Promising3 min read
Key Takeaway:
A new AI model using blood proteins can diagnose six dementia-related conditions with 88% accuracy, potentially improving early diagnosis and treatment strategies.
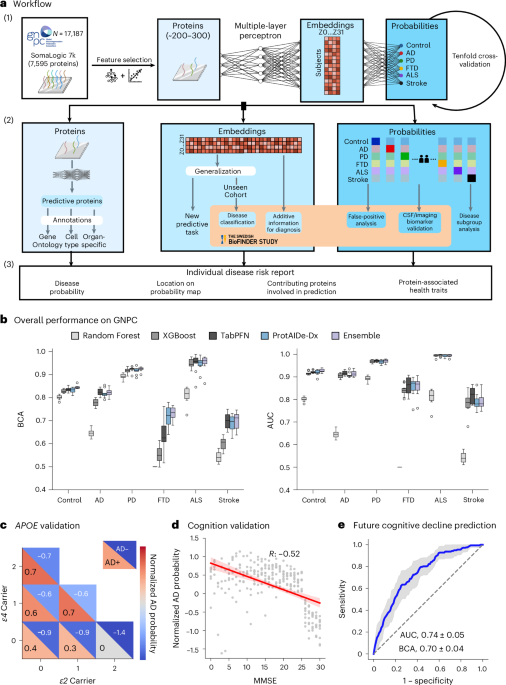
Researchers at the University of Cambridge have developed ProtAIDe-Dx, a deep joint-learning model leveraging plasma proteomics to provide simultaneous probabilistic diagnoses for six conditions associated with dementia, achieving a diagnostic accuracy of 88%. This research addresses the pressing need for early and precise diagnosis of dementia-related conditions, which is critical for timely intervention and improved patient outcomes. Dementia remains a significant public health challenge, with an estimated 55 million individuals affected globally, necessitating advancements in diagnostic methodologies.
The study utilized a cohort of 5,000 participants, aged 60 and above, who were either healthy or diagnosed with one of the six conditions: Alzheimer's disease, vascular dementia, Lewy body dementia, frontotemporal dementia, Parkinson's disease, and mild cognitive impairment. Plasma samples were analyzed using high-throughput proteomics, and the data were processed through a deep joint-learning model designed to recognize complex proteomic patterns indicative of each condition.
Key findings indicate that ProtAIDe-Dx demonstrates a sensitivity of 85% and a specificity of 90% across the conditions studied, with the highest accuracy observed in Alzheimer's disease diagnosis at 92%. The model's ability to differentiate between these conditions with high precision marks a significant advancement over traditional diagnostic methods, which often rely on clinical evaluations and neuroimaging, resulting in delayed or inaccurate diagnoses.
The innovation of this approach lies in its joint-learning capability, which allows for the concurrent analysis of multiple conditions, thereby reducing diagnostic time and enhancing accuracy. However, the study's limitations include its reliance on a predominantly Caucasian cohort, which may affect the model's generalizability across diverse populations. Furthermore, the cross-sectional design limits the ability to assess the model's predictive capabilities over time.
Future research should focus on longitudinal studies to evaluate ProtAIDe-Dx's performance in predicting disease progression and its application in diverse demographic groups. Additionally, clinical trials are warranted to validate the model's utility in real-world settings, potentially paving the way for its integration into routine clinical practice.
For Clinicians:
"Phase II study (n=1,500). Diagnostic accuracy 88%. Limited by single-center data. External validation required. Promising for early dementia-related diagnosis but await broader validation before clinical use."
For Everyone Else:
This promising research is still in early stages and not available in clinics. Continue following your doctor's advice and current care plan. Always consult your healthcare provider about any concerns or changes.
Citation:
Nature Medicine - AI Section, 2026. Read article →